Microarray: the Word to Know New Technology For New Research in Prostate Cancer Treatments
Because a new technology is directing the research activity for the discovery of “smart” drugs or “magic bullets” to fight cancer, we thought it was important to introduce this technology to our readers.
 Just five years ago, genes were studied one by one. Now, thousands are studied at one time with the development of a new technology the DNA microarray.
Just five years ago, genes were studied one by one. Now, thousands are studied at one time with the development of a new technology the DNA microarray.
A particular cell contains the entire complement of 30,000 genes and these genes send messages to convert information into active molecules (proteins) that carry on activities required to develop living systems.
But individual cells don’t use all the genes; in other words, all the genes of a cell aren’t sending messages at one time. In fact, only a small number of genes are “active” in a particular cell.
Scientists want to know which genes are active, meaning which genes are initiating messages to make proteins, and which genes are “turned off” in particular cells.
Especially, they want to know about the differences of gene activities in cancer cells and normal cells. Scientists refer to this activity as gene expression.
If they could know the gene expression in particular cancer cells, they believe they would be able to develop drugs that would interfere with the activity of those particular genes. Scientists believe they could prevent the activation of the proteins involved in developing cancer cells.
In the case of prostate cancer, one focus for scientists is finding out which genes are abnormally activated. Researchers study the activities of genes in cancerous prostate cells and compare them with the activity in normal prostate cells.
But with thousands of genes, and thousands more proteins, it would take years and years of trial and error experiments to accidentally come upon the genes affecting the development of prostate cancers.
Scientists might have an idea of what they are searching for, but, most likely, they wouldn’t be able to find it in their lifetimes unless they had better ways of looking.
“In science,” Dr. Catalona explained, “Progress comes along with the development of new tools.”
 And a new tool, the microarray, is just what researchers needed. A Stanford biochemist, Patrick Brown, developed a way to take thousands of gene molecules and put them on a single slide called a microarray.
And a new tool, the microarray, is just what researchers needed. A Stanford biochemist, Patrick Brown, developed a way to take thousands of gene molecules and put them on a single slide called a microarray.
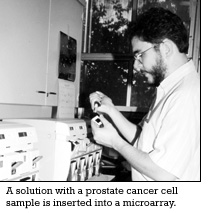
A sample of a cell being studied in the case of prostate cancer those cells would come from a prostate cancer specimen is converted into a solution that is inserted in the microarray.
A sophisticated version of a rotisserie oven “cooks” together the microarray and the solution of the prostate cancer cells being studied.
During the “baking”, the active genes from the cancer cell find their matched genes in the microarray. Only the active genes connect with their counterparts and change their visual make-up on the microarray.
These “baked goods” come out in the form of a readable chip that is fed into a laser scanner which then sends visual images to a computer screen.
At the end of the process, the image shows scientists which genes matched up in the oven. It tells them which genes are “turned on” and how much they are “turned on” in the prostate cancer cells.
The identification procedure, which could have taken years before the invention of the microarray, is now a fast process.
 This technology lets scientists know which genes are over expressed or under expressed in specific cancer cells. It opens the door to the development of drugs tailored to stop or slow the development of particular cancer cells by inhibiting the production of proteins active in those cancer cells.
This technology lets scientists know which genes are over expressed or under expressed in specific cancer cells. It opens the door to the development of drugs tailored to stop or slow the development of particular cancer cells by inhibiting the production of proteins active in those cancer cells.
If this method of analysis did no more than identify candidate genes involved in the development of prostate cancer or other cancers, it would be a revolutionary technology. But it does more.
It has the potential to be an effective diagnostic tool, identifying people susceptible to cancer and then providing essential information to prevent that cancer from developing.
Also, the same process that identifies candidate genes can be used to study the effects of drugs on those genes.
 “We’re entering into a new era in which drugs will be selected for each individual patient,” Dr. Catalona said. “In a way, it’s like the current practice of choosing different antibiotics to treat different types of infections. The model to apply for all drugs will be to look at the genetic make-up of the cancer cell and then know which drugs will be most effective for that particular genetic make-up.
“We’re entering into a new era in which drugs will be selected for each individual patient,” Dr. Catalona said. “In a way, it’s like the current practice of choosing different antibiotics to treat different types of infections. The model to apply for all drugs will be to look at the genetic make-up of the cancer cell and then know which drugs will be most effective for that particular genetic make-up.